Hi everyone,
I am currently working with ATLAS Open Data to test a Machine Learning algorithm for background estimation. I am manually streaming/downloading ROOT files rather than using the standard atlas-open-magic package.
Due to some network interruptions during the download process, I have ended up with a partial dataset. To account for these missing files, I have implemented the following workflow:
-
Luminosity Calculation
I applied the Good Runs List (GRL) to my data. To handle the missing files, I estimated an effective integrated luminosity by scaling the target luminosity by the ratio of processed events to the total expected events from the GRL. I am processing 2016 pp collision data and got an effective luminosity of approximately 25 fb⁻¹.
-
MC Normalization
For my backgrounds (Multijet Pythia8 JZ slices, Sherpa W/Z+jets, ttbar, etc.), I am using metadata that includes the cross-section ($\sigma$), number of events ($N$), and the sum of weights ($\sum w$). My scaling factor for each MC sample is cross section*L_eff/(sum of weights)
After passing both Data and MC through identical preselection cuts to define my Control Regions, the MC yields are significantly overshooting the observed Data, often by a large factor, despite the luminosity scaling. I suspect I may be mishandling the event weights. Specifically, I am looking at the branch (EventInfoAuxDyn_mcEventWeights)
I have 2 questions:
- Are there additional internal weights (like pile-up reweighting or SFs) in the Open Data branches that are mandatory to bring the MC down to the expected Data levels?
- Am I scaling the MC samples wrong?
Thank you very much
Best,
Francis
Hi @fjumawan ,
Thank you for the note, and for using the ATLAS Open Data!
When there are data transfer failures, it can be pretty tricky to ensure that the adjusted luminosity is correct. I very much recommend that you try to get as much of the data as you possibly can. The MC is much more forgiving when it comes to partial samples.
For the MC, the scaling should generally be by the cross section times the integrated luminosity times the filter efficiency times the k-factor divided by the sum of weights. For the samples you’re describing, the filter efficiencies are often below 10%, so I expect that’s the most important factor you’re missing. For the sum of weights, you can literally use the sum of the nominal weights for the sample that you’ve downloaded (instead of trying to get it from the metadata we provide) to avoid issues with the partial sample.
Best,
Zach
Good day @zmarshal,
Thank you very much for the reply. I tried your using the filter and multiplying over the sum or nominal weights. Like this
LUMI = 25.289004522843967
df_mc = df_mc.with_columns(official_scale_factor = (pl.col(“crossSection_pb”) * LUMI * 1000 * pl.col(“genFiltEff”) * pl.col(“kFactor”)) / pl.col(“raw_weights”).sum())
df_mc = df_mc.with_columns(
final_weight = pl.col("official_scale_factor")*pl.col("raw_weights")
)
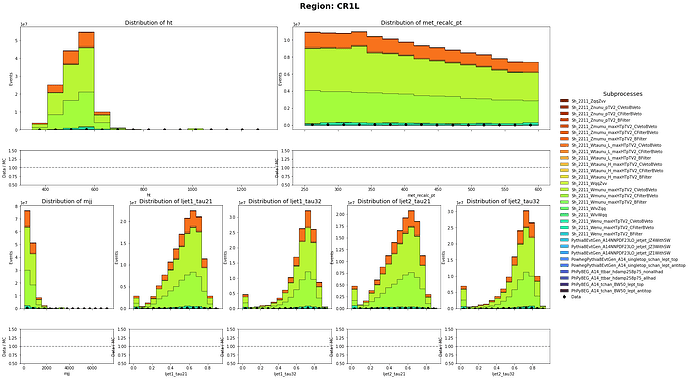
The MC samples are still to big compared to the data. For example, In my Control Region selection of 250 GeV < MET <= 600 GeV && HT <= 600 GeV && one muon and no bjets. It needs a scale of 0.00541 to match the MC and data
Before:
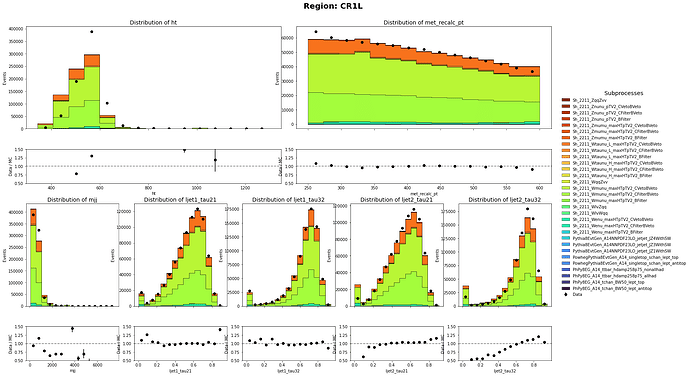
After:
Is it because I am not using the PileupWeight_NOSYS?
Best,
Francis
Hi @fjumawan ,
The pileup weight will shift the distributions around a bit, but they will not change the agreement that dramatically (it’s a few % change, typically). How are you calculating raw_weights? For the three big Wmunu samples, could you confirm what cross section and filter efficiency you apply?
Best,
Zach
Hi @zmarshal,
These are the metadata I used for the Wmunu samples
Sample: Wmunu_maxHTpTV2_BFilter
Cross Section (pb): 21806.0
genFiltEff: 0.0097968
Sample: Wmunu_maxHTpTV2_CFilterBVeto
Cross Section (pb): 21806.0
genFiltEff: 0.1460112
Sample: Wmunu_maxHTpTV2_CVetoBVeto
Cross Section (pb): 21806.0
genFiltEff: 0.8435538
Best,
Francis
Thanks @fjumawan , that looks ok to me.
What about the raw_weights column? How are you calculating it, and what do you get for the sum?
Best,
Zach
Good day @zmarshal, this is how I saved the raw_weights column from the root file
import uproot
events = uproot.open("mc20_13TeV_MC_Sh_2211_WlvWqqroot_0.root:CollectionTree").arrays(library="ak")
events["raw_weights"] = events["EventInfoAuxDyn_mcEventWeights"][:, 0]
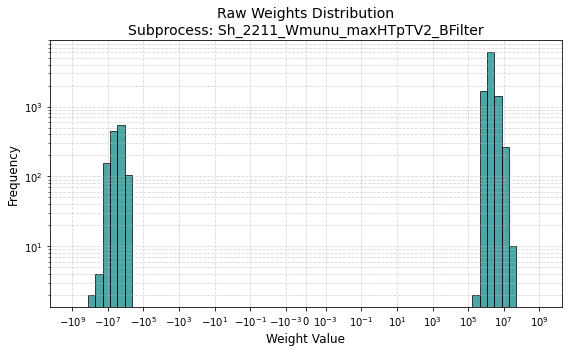
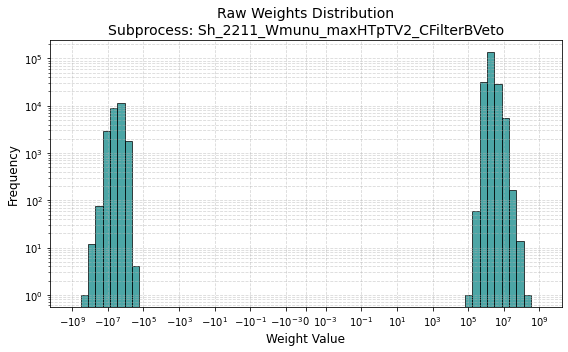
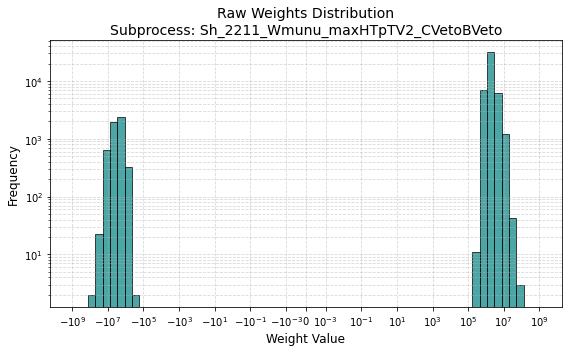
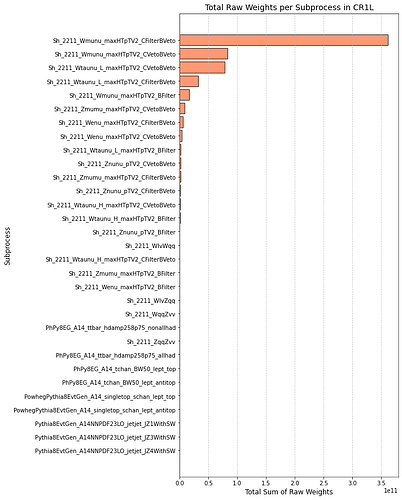
This is the distribution of weights of the region (Wmunu samples)
and here are the sum of weights
Best,
Francis
Hi @fjumawan ,
The weight distribution looks about right to me. The sum of weights looks like it’s only for a single file, though? Is that what you’re doing? Remember that the sum of weight should run over all the files in an MC sample. For the full Sh_2211_Wmunu_maxHTpTV2_BFilter sample for example, we have about 15M events, and the sum of weights is about 4.37e14.
Best,
Zach
Hi @zmarshal,
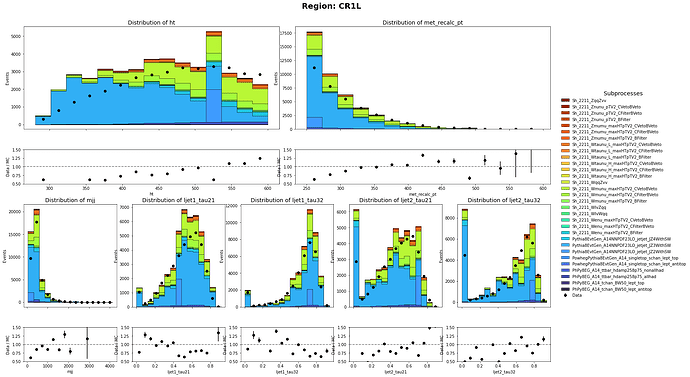
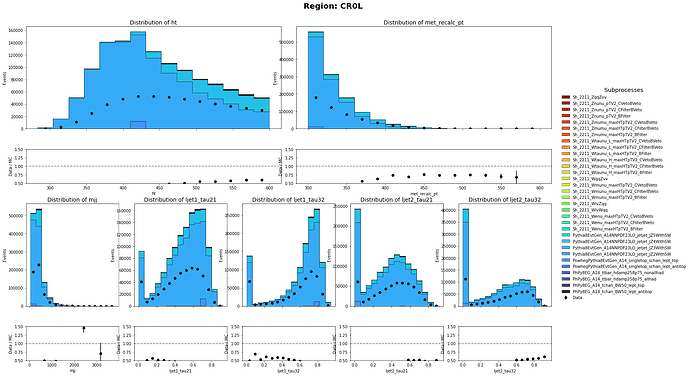
Thank you so much for pointing that out. I did only use the sum of weight per file. I have corrected it and the MC and data has a closer agreement than before. Though there are still discrepancy on the shape. Are there any problem on the weights of Multijet Samples?
CR1L definition: HT < 600 GeV, 250 GeV < MET < 600 GeV with 1 muon in the event
CR0L definition: HT < 600 GeV, 250 GeV < MET < 600 GeV with no muon in the event
Best,
Francis
Hi @fjumawan ,
Looking pretty good! I think there’s one key thing still missing: the triggers. In the real data, any event we record has to pass some trigger requirement. We have a bunch of different triggers that look for leptons, or HT, etc. The simulated samples don’t have any trigger requirements. When you compare the data and simulation, you need to make sure you’re comparing events that pass the same trigger. We have a few different branches for that in the E&O open data, depending on which trigger you want to use.
https://opendata.atlas.cern/docs/data/for_education/13TeV25_details
Best,
Zach
Hi @zmarshal,
Thank you so much for the guidance!
I am currently using the 2024 Data for research version. So I am accessing the physlite directly and it does not have the branches described in the “https://opendata.atlas.cern/docs/data/for_education/13TeV25_details”
Is there an equivalent branches present in the physlite? I am trying to find these triggers
HLT_xe90_mht_L1XE50, HLT_xe100_mht_L1XE50, HLT_xe110_mht_L1XE50
which are not present in the AnalysisTrigMatch branches found in this documentation: https://atlas-physlite-content-opendata.web.cern.ch/
Is it possible to apply scaling instead to imitate the effects of trigger cuts?
Best,
Francis
Hi @fjumawan ,
You can see how we’re doing things here:
https://gitlab.cern.ch/atlas-outreach-data-tools/physlitetoopendata/-/blob/main/Root/PhysLiteToOpenData.cxx?ref_type=heads#L264
If you aren’t running some kind of C++ framework around this (if you’re just reading branches and drawing), then this is tricky to emulate. The easiest thing is to ensure that you are well above the threshold for the relevant trigger, for example by requiring 250 GeV of MET if you’re interested in MET triggers. That should be relatively safe. (There are some extra effects because the data and MC are not being selected only with that trigger, but I hope those will turn out to be small-ish)
Best,
Zach
1 Like